IMAT Enhances Clinical Trial Cohort Identification
IMAT Enhances Clinical Trial Cohort Identification
Introduction
 Timely access to data is always a top priority for mature organizations. Identifying and acting on the information housed in an organization’s data repositories can literally mean the survival of a company. The problems associated with connecting with this data, however, are many and varied. In clinical trial research, for example, organizations need to index and query databases with billions of rows of data representing many millions of unique patients. Queries designed to identify clinical trial cohorts are often complex and computationally intensive. They may take multiple hours to days to run, restricting what can be done during the research period and limiting the research questions that can be asked.
Timely access to data is always a top priority for mature organizations. Identifying and acting on the information housed in an organization’s data repositories can literally mean the survival of a company. The problems associated with connecting with this data, however, are many and varied. In clinical trial research, for example, organizations need to index and query databases with billions of rows of data representing many millions of unique patients. Queries designed to identify clinical trial cohorts are often complex and computationally intensive. They may take multiple hours to days to run, restricting what can be done during the research period and limiting the research questions that can be asked.
IMAT Solutions has developed a unique indexing and query engine that can combine relevance ranked, full text, Boolean, meta data, and field searches including text, date/time, and numeric variables. All this can be done in the same indexing structure to dramatically increase query speeds and enhance accuracy while significantly reducing infrastructure requirements and server load. This summary describes a collaborative study that used IMAT with its specialized indexing and query tools to mimic a recent clinical trial. Remarkably, IMAT produced a reduction in cohort selection time to seconds versus the hours or even days as is typical of existing solutions.
Advantages of the IMAT Architecture
We were working with a company that is using IMAT in their medical analytics system. A particular query had generated 3,500 additional queries that would need to be run. The queries were being executed by IMAT in under a minute and we were apologetic about the “slow” speed. “That’s nothing,” they said, “it takes a day to process these queries using our old approach using a database.”
IMAT has received eleven patents on its unique approach to search. Every stage of the search process, from a pipeline architecture indexing phase to ranked, Boolean or ranked OR search, has been optimized for speed and minimal CPU usage. As an example of this optimization, if a Boolean AND search is done and one term has 50 million hits and another has 400 hits, the search speed will be most related to the 400 hit term rather than the 50 million hit term.
A New Kind of Search Index
Another key difference in the IMAT approach is the number and variety of terms that are in the index. A typical traditional search system has millions of unique terms that can be searched, ordered in some kind of b-tree or other structure. IMAT creates billions of unique patterns related to subsets and combinations of possible search terms along with field and category variables.
Full text, numeric, and date/time patterns are all indexed in the same index structures. The access methods for this index are accelerated by structures that use some of the fastest internal computer CPU and memory operations possible. Indexes can be created or updated as fast or faster than other search systems.
A New Approach to Relevance Ranking Computations
In a traditional search system’s relevance ranked search, each potential hit, record, or document, is processed with relevance boosts at every potential word, record, or document level. The very last hit of the very last term might end up being the highest ranked using this “bubble up” approach to relevance.
Instead of this CPU intensive process of relevance ranking, IMAT uses micro searches to carefully hone in on the highest ranked hits first, then the next highest ranked, and so forth until enough hits have been retrieved, often without even considering the areas of hits that are in the lower levels of relevance. This accelerated approach is also used to speed up Boolean and weighted searches. The end result is an order of magnitude or more speed advantage for the search process while being able to equal or approximate different relevance weightings and strategies used by the traditional search or database engines.
Multiple Gradations of Meaning in Search
The searching process involves separating out various sets and subsets of hits that can be complex when the search tries to distinguish multiple semantic distinctions. Ontologies and classification systems used in a search can trigger many related searches as various possible avenues are explored during the search. This approach, taken to its logical conclusion, can end up with large complex queries involving thousands of terms. Once again, a traditional system can be overwhelmed by such a search and become extremely slow.
IMAT Architecture Matches Solid State Drive Capability
One of the exciting new developments in recent years has been solid state drive technology, such as the FusionIO drive. These drives provide a tremendous speed boost for a process that involves many seeks to positions in a file and small reads. If a search involves retrieving millions of bytes from the disk with extensive processing, as occurs in a standard search system, there is often little advantage to have the solid state drive over a fast spinning disk system.
IMAT is designed to find exactly what it needs to process in the indexes, seek to specific locations, and read and rapidly process a few bytes from the disk record. A great example of an apples to apples search query comparing IMAT and another well known system ended up having IMAT process 500 kilobytes while the other system processed 500 megabytes. The IMAT portion consisted of many seeks within the index files. Five hundred seeks per second on disk might be replaced by 100,000 seeks per second on the solid state drive. IMAT’s speed can sometimes jump 20 to 100 times faster on the solid state drive over the traditional spinning drive.
Health Care Database
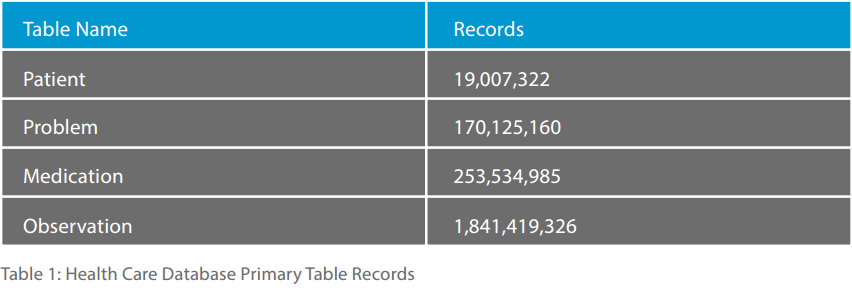
The health care database used for the study consisted of over 2 billion records for over 19 million patients with medical problem (or diagnosis), medication and observation (or lab test or other diagnostic reading) table groups. Of the 19 separate tables, we fed records to IMAT using the patient table, the two problem tables, the two medication tables, and the two observation tables. The records were ingested using multi threaded feeds into an IMAT system version 4.1 for Linux on a 24-core, 64G RAM, 10k rpm raided drive system. No solid state drives were used in this study. The total elapsed time for ingestion of the data was about 2.5 days.
Table 1 shows the size of each of these four information tables totaling 2.3 billion records.
Cohort Selection Study Searches
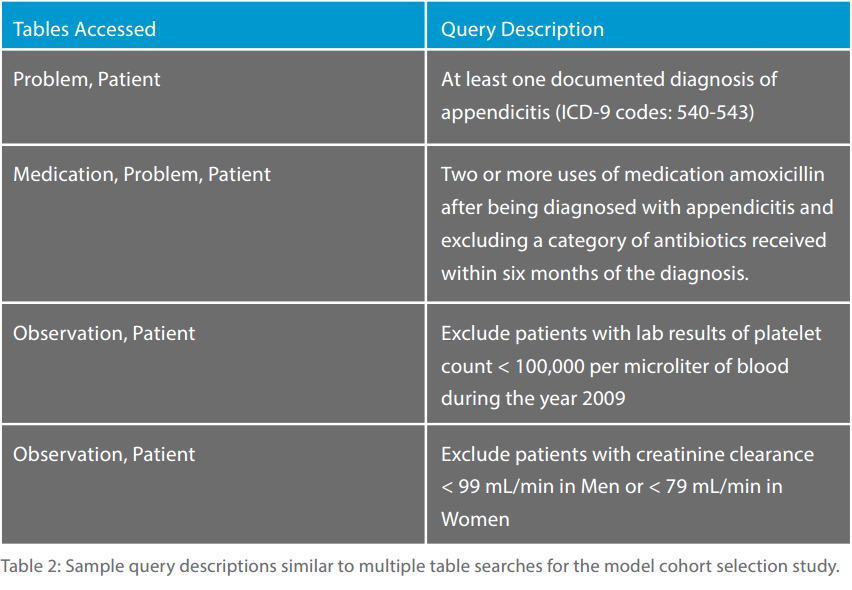
The sample cohort selection study was simulated by selecting sets of possible patients via the ICD-9 medical problem classification code groups. Afterwards, the procedure would intersect or exclude another set of patients from another condition. Conditions include specific medications or categories of medications, additional diagnostic code ranges, value ranges for observation tests that were performed and date conditions such as the treatment medication that was taken after the diagnosis was given.
Table 2 shows sample query descriptions that were similar to those for this cohort study.
IMAT Job Step Processing
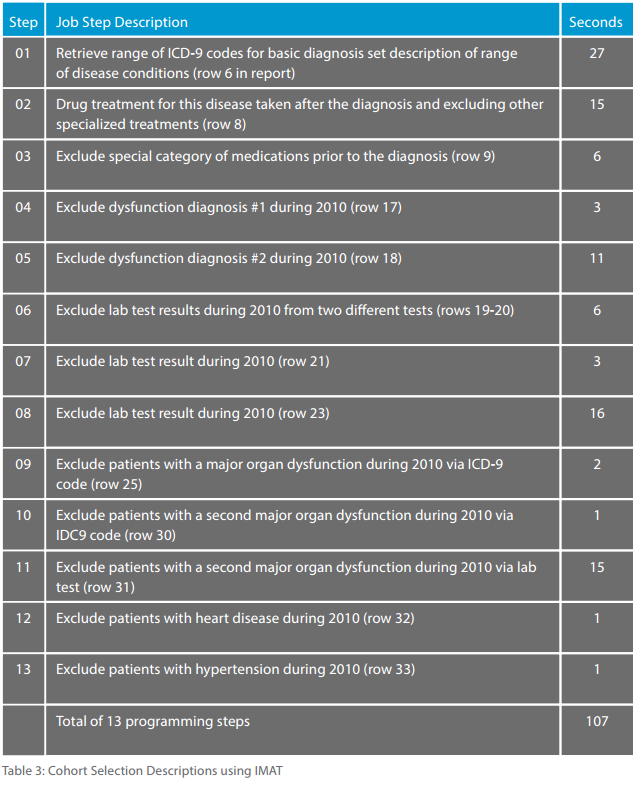
This IMAT-based implementation closely paralleled the cohort study with 13 separate procedural steps. Using the same IMAT solution previously described, the 13 steps were processed as a separate Cohort Selection module that uses multiple calls to the IMAT system. Results on individual searches ranged from hundreds of millions of records for one search to only 14 records for another. There were many levels in between such as 130,000 qualifying records for one exclusion search. Table 3 shows a very general description of the 13 steps and the time in seconds for each step and complete search sequence. The entire search procedure took only 107 seconds.
Job Step XML Specification
The individual job steps and query results needed to create a cohort set are relatively easy to create for users with some degree of database skills. The basic programming unit of a “job step” is described as an initial set specification or a Boolean operation (AND, OR or binary AND NOT). The Boolean operation indicates this job step’s interaction with the cumulative set that is being created for the cohort. The query section uses Boolean operators and relatively simple search syntax, and also may include other more complex specifications such as time order relationships between matching rows in multiple tables. The common Boolean space is specified in a key name and the output xml in the Boolean space is output to a file containing the problem, medication or observation space and another file containing the patient space search results. These files containing result sets can then be combined via other Boolean operations to further refine the final set until it ends up as the patient list recommended for the cohort.
Figure 1 shows a sample job set xml specification.

Conclusion
Complex searches of large databases, while critical, often require overnight and weekend runs as the complex search criteria uses many records in each table. IMAT is able to ingest these databases quickly and evaluate complex query sequences in near real time. The ability to retrieve and evaluate results in minutes or even seconds and to then try a wide variety of queries would transform the search tool into an extension of the researcher’s mind, creating the possibility of performing hundreds of times the amount of searches than were previously possible. More “what if?” exploration can be performed. More avenues and dead ends can be explored without draining precious time and resources.
About IMAT Solutions
IMAT Solutions is a new, disruptive medical data management and decision support company. Every day, our software processes tens of millions of records to help 31,000 healthcare professionals working for 2,000 Health Information Exchanges, Accountable Care Organizations, Medical Homes, hospitals and clinics to provide the best possible healthcare to almost 20 million patients. Based on patented technology, IMAT has been developed to address the ever more complex and demanding needs of today’s healthcare providers and practitioners. Our product is based on a completely new approach to ingesting, storing and processing massive amounts of structured and unstructured data from EHRs, lab results, physician notes, transcripts, ADTs and letters. This patented design delivers disruptive speed, superior data accuracy, easy to use charting, reports and queries and is highly cost-effective.


